EvoRator Gallery
Click on the linked example titles to view EvoRator results and the rotatable molecules colored according to EvoRator conservations scores using ConSurf coloring scheme, or click on the static figures to see them in greater detail.
Color coding scheme of EvoRator/ConSurf
Example 1: Dermcidin (PDB ID 2YMK chain A).
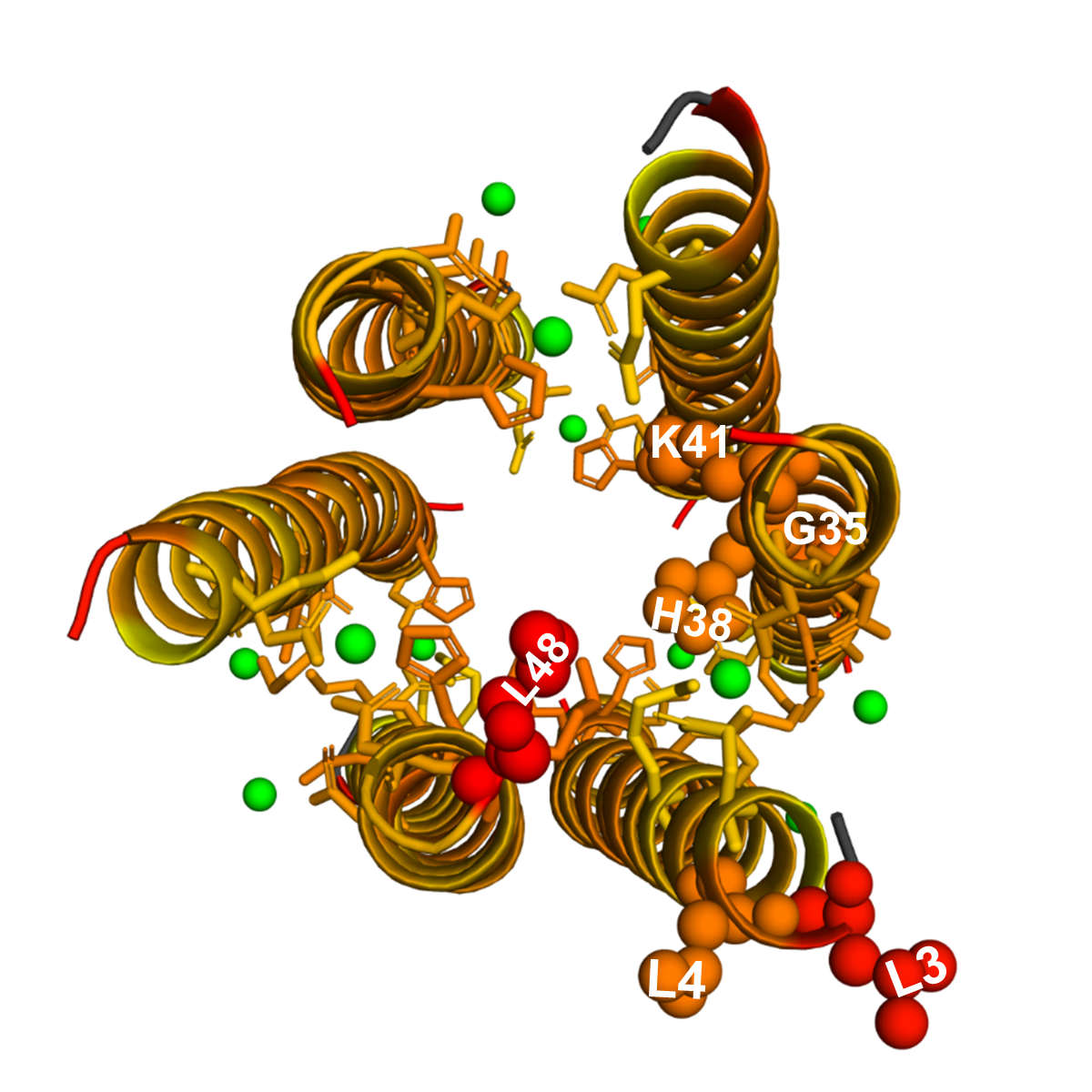
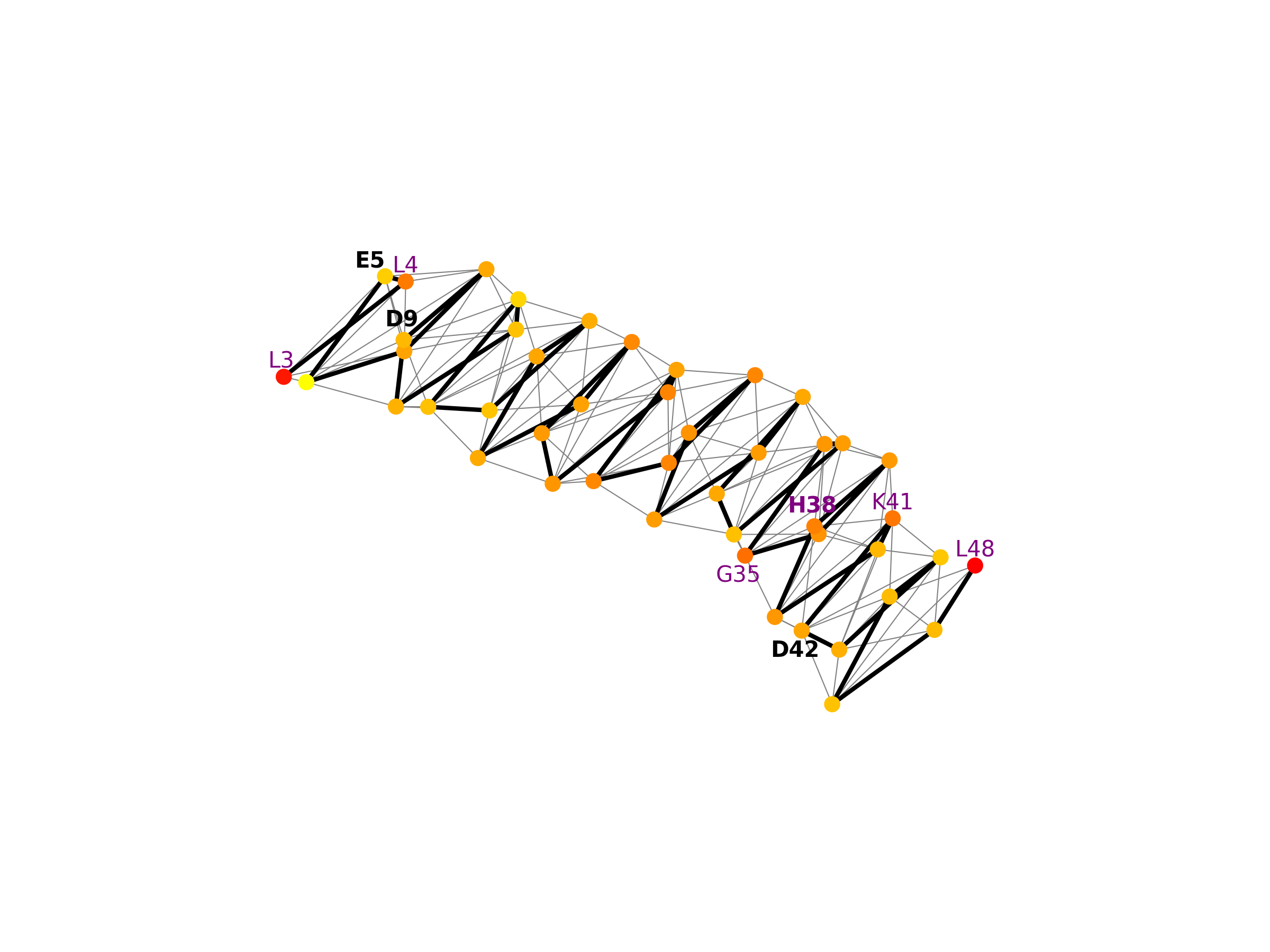
Orphans are proteins with few or no homologs in other species. ConSurf cannot reliably estimate residue-level evolutionary rates for orphan proteins, because it requires multiple sequence alignment of large number of homologous sequences. Dermcidin (Fig. dermcidin_1) is an orphan antimicrobial peptide that forms ion channels that are composed of zinc-connected trimers of antiparallel helix pairs in the membranes of bacterial cells. Zinc binding involves coordination that is not easily achieved, and imposes very strong preference for specific polar residues. Therefore, residues H38, D42, E5, and D9, which bind zinc, are expected to be highly conserved. EvoRator predicts that most neighboring residues of H38, D42 and E5 should be highly conserved (Fig. dermcidin_2). L3 and L4, which are neighbors of E5, are ranked 2nd and 5th most conserved residues. G35, which is a neighbor of H38, is ranked 3rd most conserved residues. K41, which is a neighbor of both H38 and D42, is ranked 4th most conserved residues. H38, which binds zinc, is the 6th most conserved residue. Here we show the structure marked by an alternative coloring scheme that unlike ConSurf, uses the raw predicted values, which in the case of dermcidin better reflects the differences in conservation scores between residues. You can download a high resolution figure for pymol from here.
 |
 |
| Fig. dermcidin_1 | Fig. dermcidin_2 |
Example 2: Endolysin (PDB ID 2LZM chain A).
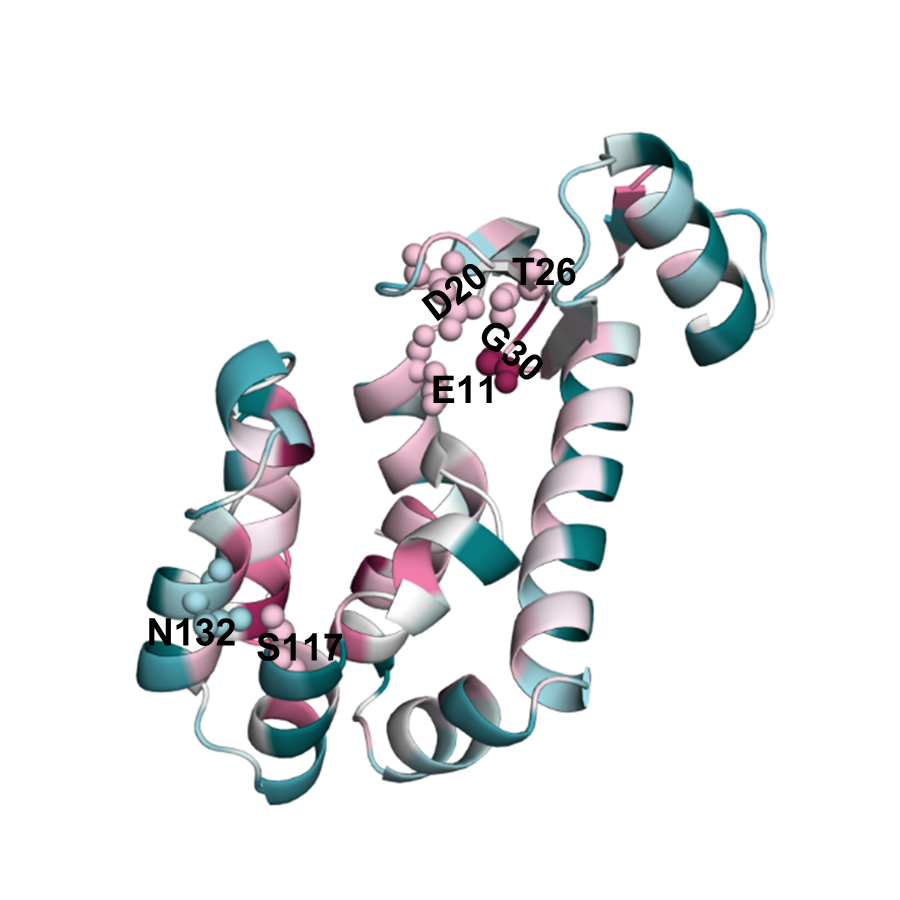
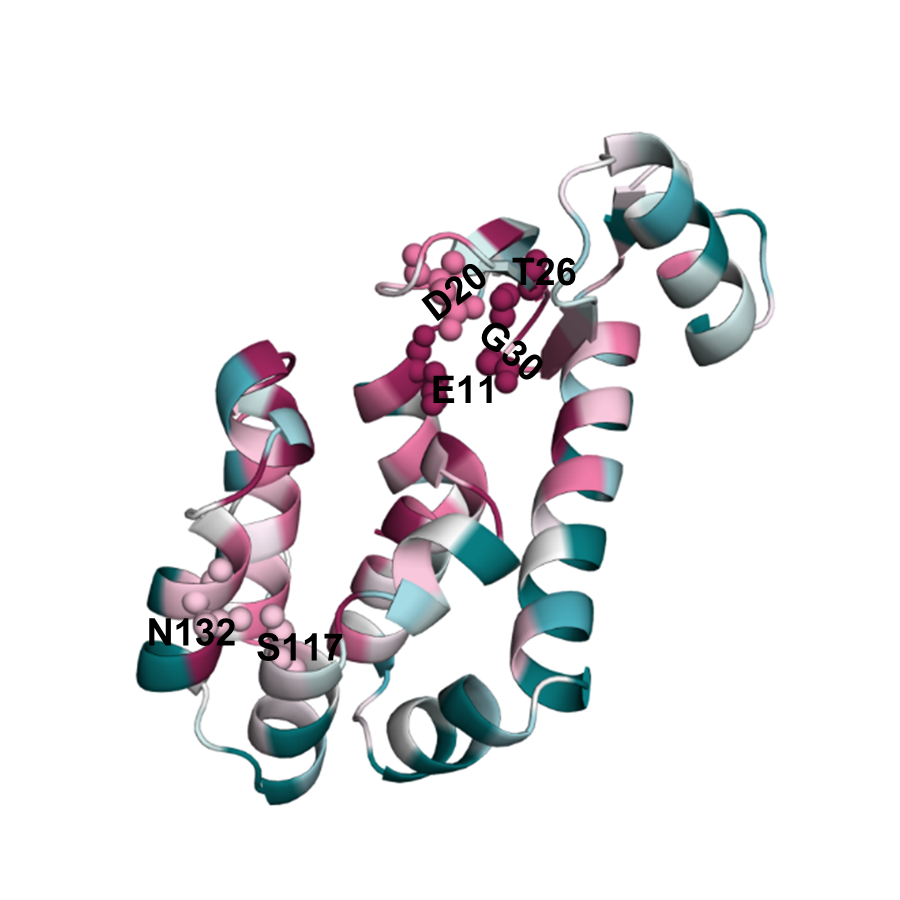
Bacteriophage T4 endolysin is an enzyme that degrades the cell wall of bacterial hosts. Endolysin has six residues that when mutated lead to loss of its catalytic activity. Therefore, these residues are expected to be highly conserved due to purifying selection. EvoRator prediction (Fig. endolysin_evorator) is highly similar to the prediction of ConSurf (Fig. endolysin_consurf), correctly indicating the conserved functional patches created by these six residues.
 |
 |
| Fig. endolysin_evorator | Fig. endolysin_consurf |
Example 3: SARS-CoV-2 spike glycoprotein ectodomain (PDB ID 6VYB chain A).
The evolutionary rates estimated by ConSurf can be unreliable when there is too little or too much sequence variation among homologs (see here and here for details). EvoRator can predict the rates of such residues using structural information as well as by taking into account the reliable conservation scores of neighboring positions. Shown here is the prediction of ConSurf (see here). for spike glycoprotein augmented by the prediction of EvoRator (Fig. spike_evorator). Sites that were previously assigned with unreliable rates by ConSurf (light yellow) are are instead marked by EvoRator prediction. Highlighted are six such sites at which mutations with functional implications occurred: H69 and V143 were deleted whereas S373, K417, Y505, and D614 were substituted in SARS-CoV-2 variant Omicron compared with the ancestral Wuhan-Hu-1 spike. EvoRator predicts that sites at which deletions occurred are the most variable ones among 157 sites that were mapped onto the 3D structure and assigned with unreliable rates by ConSurf. These deletions were shown to impact viral infectivity and antigenicity (see GISAID).
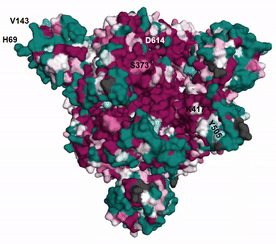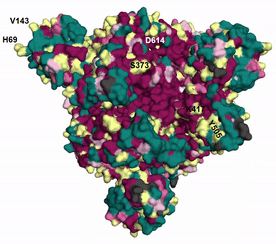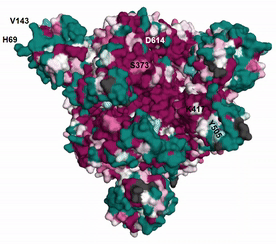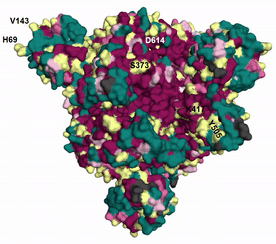 |
| Fig. spike_evorator |